Phospholipase A2 (PLA2) enzymes are known to play a role in many inflammatory diseases, including asthma, arthritis and atherosclerosis. It then stands to reason that PLA2 inhibitors could represent a new class of anti-inflammatory medication. To better understand PLA2 enzymes and help drive therapeutic drug development, researchers at University of California, San Diego School of Medicine developed 3D computer models that show exactly how two PLA2enzymes extract their substrates from cellular membranes. The new tool is described in a paper published online the week of Jan. 26 by the Proceedings of the National Academy of Sciences.
“This is the first time experimental data and supercomputing technology have been used to visualize an enzyme interacting with a membrane,” said Edward A. Dennis, PhD, distinguished professor of pharmacology, chemistry and biochemistry and senior author of the study. “In doing so, we discovered that binding the membrane triggers a conformational change in PLA2 enzymes and activates them. We also saw several important differences between the two PLA2 enzymes we studied — findings that could influence the design and development of specific PLA2 inhibitor drugs for each enzyme.”
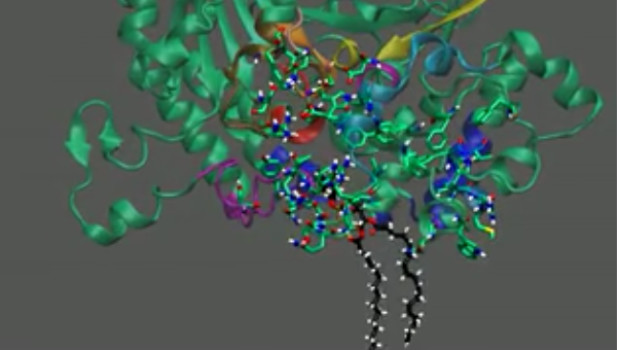
First 3D computer model of phospholipass A2 enzyme extracting its substrate from cell membrane.
The computer simulations of PLA2 enzymes developed by Dennis and his team, including first author Varnavas D. Mouchlis, PhD, show the specific molecular interactions between PLA2 enzymes and their substrate, arachidonic acid, as the enzymes suck it up from cellular membranes.
Make no mistake, though — the animations of PLA2 in action are not mere cartoons. They are sophisticated molecular dynamics simulations based upon previously published deuterium exchange mass spectrometry (DXMS) data on PLA2. DXMS is an experimental laboratory technique that provides molecular information about the interactions of these enzymes with membranes.
“The combination of rigorous experimental data and in silico [computer] models is a very powerful tool — the experimental data guided the development of accurate 3D models, demonstrating that these two scientific fields can inform one another,” Mouchlis said.
The liberation of arachidonic acid by PLA2 enzymes, as shown in these simulations, sets off a cascade of molecular events that result in inflammation. Aspirin and many other anti-inflammatory drugs work by inhibiting enzymes in this cascade that rely on PLA2 enzymes to provide them with arachidonic acid. That means PLA2 enzymes could potentially also be targeted to dampen inflammation at an earlier point in the process.
Story Source:
The above story is based on materials provided by University of California – San Diego. The original article was written by Heather Buschman. Note: Materials may be edited for content and length.
Journal Reference:
- Varnavas D. Mouchlis, Denis Bucher, J. Andrew Mccammon, and Edward A. Dennis. Membranes serve as allosteric activators of phospholipase A2, enabling it to extract, bind, and hydrolyze phospholipid substrates. PNAS, January 2015 DOI: 10.1073/pnas.1424651112